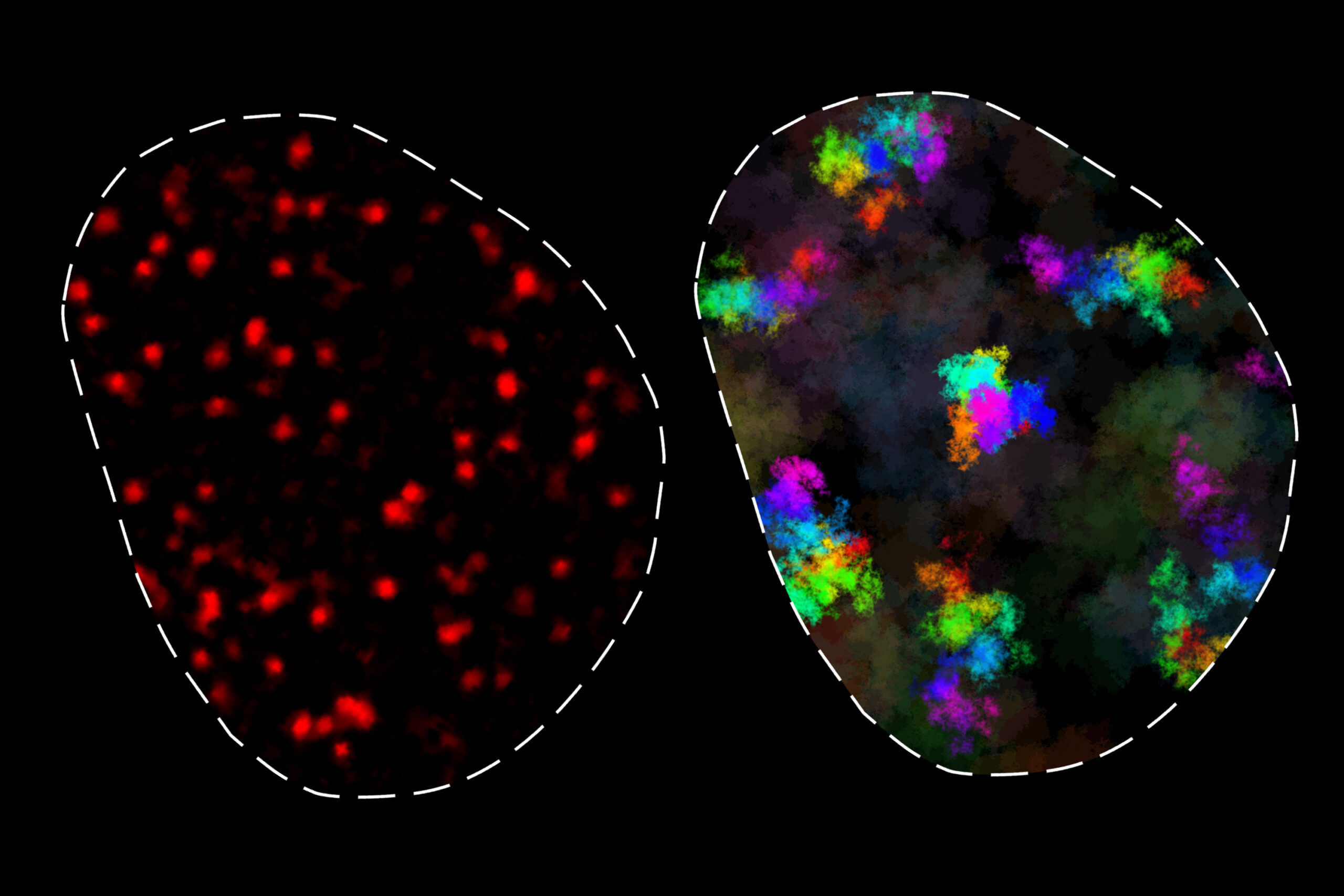
The control of gene expression partially relies on interactions between genes and regulatory elements along the genome. These interactions are influenced by chromatin, a DNA-protein mix, which moves within a crowded environment. MIT researchers have, for the first time, quantified chromatin movement over timescales from microseconds to hours.
Their study shows chromatin exists in two states: one where it moves in a limited manner, mainly contacting nearby genome regions, and another where it moves more freely, reaching distant regions over longer times. These observations provide insights into gene regulation and chromatin’s role in processes like DNA repair.
“We could observe chromatin motion over an unprecedented range,” stated Anders Sejr Hansen, MIT biological engineering professor and senior study author, published in Nature Structural and Molecular Biology. Lead authors include MIT postdoc Matteo Mazzocca, PhD students Domenic Narducci and Simon Grosse-Holz, with contributions from Jessica Matthias and Tatiana Karpova.
Chromatin is often depicted as static, but it constantly moves to enable gene interactions with distant DNA regulatory sequences and facilitate DNA repair. “Chromatin dynamics are crucial for all nuclear processes,” Hansen noted. The movement of specific genome locations is constrained by DNA’s polymer nature, which pulls back any movement.
This behavior, termed subdiffusive movement, has been inconsistently reported due to previous studies’ inability to track long-term movements. The MIT team employed MINFLUX, a super-resolution microscopy technique developed by Nobel laureate Stefan Hell, to track tiny object movements over extended periods.
“MINFLUX bypassed conventional microscopy limitations, allowing faster and longer chromatin movement measurements,” said Narducci. The researchers analyzed cells over timescales from 200 microseconds to 10 seconds, extending to several hours by combining imaging techniques.
The research identified two chromatin dynamics classes across various mouse and human cell types. Over short to intermediate timescales, loci typically move within about 200 nanometers, indicating a stronger subdiffusive pull than previously thought. “There’s a region of influence for genomic loci,” Grosse-Holz explained.
This proximity aids DNA repair by keeping broken strands close. Genes and regulatory elements within 100,000 base pairs find each other without extra assistance, as Mazzocca explained. The second dynamic class allows chromatin to move over greater distances, but only over longer timescales, differing across cell types.
“The behavior varies significantly between cell types,” Hansen remarked, noting existing chromatin dynamic models like the Rouse and fractal globule models don’t fully explain the findings. The study suggests incorporating factors like chromatin-nucleoplasm interactions. “These findings confirm chromatin’s subdiffusive motion and its consistency across cell types and timescales,” said Luca Giorgetti of the Friedrich Miescher Institute.
The research received support from the National Institutes of Health, the National Science Foundation CAREER Award, a Pew-Stewart Scholar for Cancer Research Award, and the Bridge Project, a collaboration between MIT’s Koch Institute and the Dana-Farber/Harvard Cancer Center.
Original Source: news.mit.edu